
Features
Seed & Chemical
Seed Treatment
Rotate clubroot resistant varieties
As many growers know, resistant varieties are a critical tool for managing clubroot, a devastating canola disease. So Alberta researchers are assessing the pathogen’s ability to erode the effectiveness of the resistance genes and developing guidance to help growers protect this valuable tool.
Clubroot is caused by the soil-borne pathogen Plasmodiophora brassicae. It infects the roots of crops in the Brassica genus, such as canola and cole crops. Galls form on the roots, preventing water and nutrients from getting to the rest of the plant. When the galls decay, they leave millions of resting spores in the soil. Those spores can survive for up to about 20 years, waiting and ready to infect the next Brassica host that grows in the field.
Although the existing clubroot-resistant canolas are currently working very well, the researchers were concerned about the pathogen’s ability to fight back.
“We knew generally from other pathogens and their hosts that pathogens can evolve in response to the selection pressure that is imposed by growing resistant varieties of the host crops. And specifically, we knew of instances involving clubroot, for instance with oilseed rape in Europe and cruciferous vegetables in the U.S., where the resistance had eroded or in some cases broken down after repeated cropping of a resistant variety in a particular piece of land,” explains Dr. Stephen Strelkov of the University of Alberta.
So Strelkov and his lab conducted two greenhouse experiments to investigate this issue, collaborating with Dr. Kelly Turkington of Agriculture and Agri-Food Canada (AAFC) and Dr. Sheau-Fang Hwang of Alberta Agriculture and Rural Development (ARD) on these and other ongoing experiments.
“In the first experiment, we wanted to assess the effect of multiple infection cycles on the clubroot pathogen’s ability to cause disease. So we took a population and a single-spore isolate of pathotype 3 and inoculated a soil potting mix,” says Strelkov. Clubroot pathotypes, or strains, are identified based on their ability to cause disease in a standardized set of hosts. Pathotypes in Alberta include pathotypes 2, 3, 5, 6 and 8. Pathotype 3, a very virulent type, is the most common pathotype in canola crops in the province.
In the inoculated potting mix, the researchers planted a selection of resistant (R), moderately resistant (MR) and susceptible (S) hosts. These included: three Argentine canola cultivars (Brassica napus) (R, MR and S), two Polish canola cultivars (Brassica rapa) (R and R), a kale cultivar (Brassica oleracea) (MR), and a Chinese cabbage cultivar (Brassica rapa) (S).
They grew 16 seedlings of each host, with one seedling per pot, in a replicated experiment. After six weeks, they pulled the plants out of the soil and checked the roots for clubroot galls. They rated the disease level in the roots, and if there were any galls, they harvested the spores and placed them back in the same pot in the same soil. Then they grew a new set of 16 seedlings from the same host in the same pots with the same soil. They repeated this process for five cycles for each host.
In some of the cultivars, the disease level didn’t change very much from cycle to cycle. But in other cultivars, the resistance was significantly eroded (see Fig. 1).
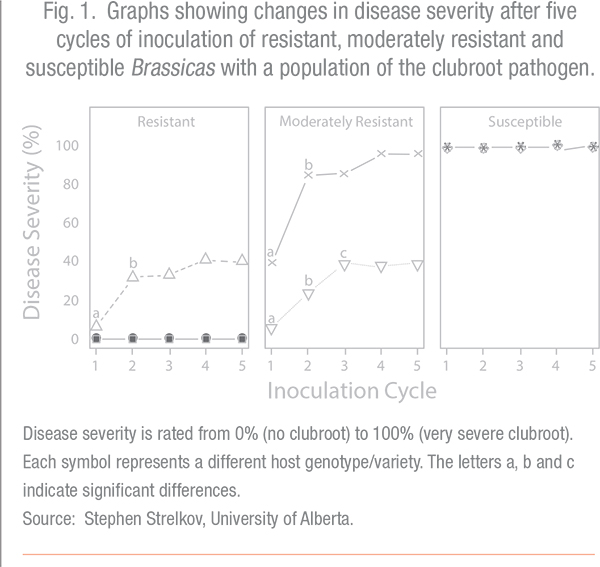 |
Strelkov notes, “In some cases, we found that repeated cropping could erode the effectiveness of the resistance. For instance some varieties that had been moderately resistant became susceptible or highly susceptible. And it usually didn’t take very long – maybe one or two cycles; so they went from a disease severity of around 40 per cent to about 90 or 100 per cent after a couple of cycles. And a [Brassica napus] resistant cultivar went from a disease severity of about five or six per cent to about 40 per cent.”
The main aim of the second experiment was to see whether the commercial canola cultivars grown on the Prairies carry the same or different sources of resistance. The particular set of genes a company uses for clubroot resistance in its cultivars is not public information; it’s the company’s proprietary information. But knowing whether or not cultivars have different genes for resistance could help growers to manage clubroot by rotating cultivars with different resistance sources.
For this experiment, the researchers harvested the pathogen populations that had been cycled with the cultivars in the first experiment. They used those populations to inoculate four resistant canola cultivars from two other companies. And then they evaluated the level of disease in those four cultivars.
“If two cultivars have the same source of resistance, then the pathogen populations that are cycled in one cultivar should show increased infectivity on the other cultivar,” explains Strelkov. “But for the cultivars we tested, the pathogens weren’t cross-infective. It’s preliminary data, but maybe, hopefully, there are at least some different sources of resistance out there.”
Another positive sign from these experiments is that the resistance in the two Polish canola cultivars was not eroded over the five cycles. Strelkov says, “That could be material for breeders to try to see if they could incorporate that resistance into Brassica napus types because [the Brassica rapa types] certainly seemed to withstand infection more than the Brassica napus cultivars we tested.”
Strelkov and his colleagues are continuing their studies on clubroot resistance. One of their current projects is using another approach to explore whether the different canola cultivars may have some different resistance genes.
“We’re finding that some resistant cultivars appear to have a different profile of how the pathogen proliferates inside [the roots], not only how much it proliferates but also when it proliferates and so on,” he explains. “So we’re comparing [the pathogen proliferation profiles] in the different resistant types and combining that with [examination of the tissues] to look at the infection process. This could provide information as to whether the same resistance mechanisms may be at play or if different ones may be involved.”
Implications for canola production
“In our experiment, we saw that growing the same resistance source twice in a row under high disease pressure already produced some pathogen strains that were causing high levels of the disease. So I think it wouldn’t take very long for this to happen under field conditions if a farmer were continuously growing the same resistance source back to back for maybe three years in a row,” notes Strelkov.
“That’s what was seen in the U.S. with vegetables and in the U.K. [with oilseed rape]. When they grew the same resistance source for about three years in a row, they started to see increased disease levels.”
According to Strelkov, the best option for resistance stewardship is crop rotation. In particular, growers should try to avoid canola-on-canola and very short canola rotations.
As an additional tool to protect resistance, rotating resistant cultivars from the different companies may help. “If you always grow the same cultivar, then you know for sure that it carries the same resistance genes. But if you grow different cultivars, at least there is some potential that there are some different resistance genes,” he says.
Strelkov emphasizes, “Clubroot management is all built around deployment of genetically resistant cultivars. If we abuse the resistance by cropping in extremely short rotations, over and over, then the resistance could become less effective. The companies of course will try to find other sources of resistance but it will become harder.
“Resistance is such an important tool, and we don’t want to lose it. Taking care of the resistance [through crop rotation] is crucial to keep our main tool for managing clubroot.”
April 9, 2014 By Carolyn King
 Resistant canola varieties are a critical tool for managing clubroot. As many growers know
Resistant canola varieties are a critical tool for managing clubroot. As many growers know