
Features
Agronomy
Genetics/Traits
Plant geneticists develop a new application of CRISPR to break yield barriers in crops
Scientists at Cold Spring Harbor Laboratory (CSHL) have harnessed the still untapped power of genome editing to improve agricultural crops. Using tomato as an example, they have mobilized CRISPR/Cas9 technology to rapidly generate variants of the plant that display a broad continuum of three separate, agriculturally important traits: fruit size, branching architecture and overall plant shape.
All are major components in determining how much a plant will yield. The method is designed to work in all food, feed, and fuel crops, including the staples rice, maize, sorghum and wheat.
"Current rates of crop yield increases won't meet the planet's future agricultural demands as the human population grows," says CSHL Professor Zachary Lippman, who led the research. "One of the most severe limitations is that nature hasn't provided enough genetic variation for breeders to work with, especially for the major yield traits that can involve dozens of genes. Our lab has now used CRISPR technology to generate novel genetic variation that can accelerate crop improvement while making its outcomes more predictable."
The team's experiments, published in Cell, involve using CRISPR to make multiple cuts within three tomato genome sequences known as a promoters -- areas of DNA near associated genes which help regulate when, where, and at what level these "yield" genes are active during growth. In this way generating multiple sets of mutations within each of these regulatory regions, the scientists were able to induce a wide range of changes in each of the three targeted traits.
"What we demonstrated with each of the traits," explains Lippman, "was the ability to use CRISPR to generate new genetic and trait variation that breeders can use to tailor a plant to suit conditions. Each trait can now be controlled in the way a dimmer switch controls a light bulb."
By using CRISPR to mutate regulatory sequences -- the promoters of relevant "yield" genes rather than the genes themselves - the CSHL team finds that they can achieve a much subtler impact on quantitative traits.
Fine-tuning gene expression rather than deleting or inactivating the proteins they encode is most likely to benefit commercial agriculture because of the flexibility such genetic variation provides for improving yield traits.
"Traditional breeding involves great time and effort to adapt beneficial variants of relevant genes to the best varieties, which must continuously be improved every year," says Lippman. "Our approach can help bypass this constraint by directly generating and selecting for the most desirable variants controlling gene activity in the context of other natural mutations that benefit breeding. We can now work with the native DNA and enhance what nature has provided, which we believe can help break yield barriers."
Each of the mutated areas creates what are known as quantitative trait loci (QTL). In any given plant, QTL have arisen naturally over thousands of years, the result of spontaneous mutations that caused subtle changes in yield traits.
Searching for and exploiting QTL from nature has been an objective of plant breeders for centuries, but the most valuable QTL - those that cause subtle changes in traits - are rare. Lippman and his team have now shown that CRISPR-generated QTL can be combined with existing QTL to create "toolkits" of genetic variation that exceed what is found in nature.
The research discussed here was supported by a PEW Latin American Fellowship; a National Science Foundation Postdoctoral Research Fellowship in Biology grant (IOS- 1523423); a National Science Foundation Plant Genome Research Program grant (IOS-1732253); and a National Science Foundation Plant Genome Research Program grant (IOS-1546837).
"Engineering quantitative trait variation for crop improvement by genome editing" appears online in Cell September 14, 2017. The authors are: Daniel Rodríguez-Leal, Zachary H. Lemmon, Jarrett Man, Madelaine E. Bartlett, and Zachary B. Lippman. The paper can be viewed at: http://www.cell.com/cell/newarticles
September 29, 2017 By Cold Spring Harbor Laboratory
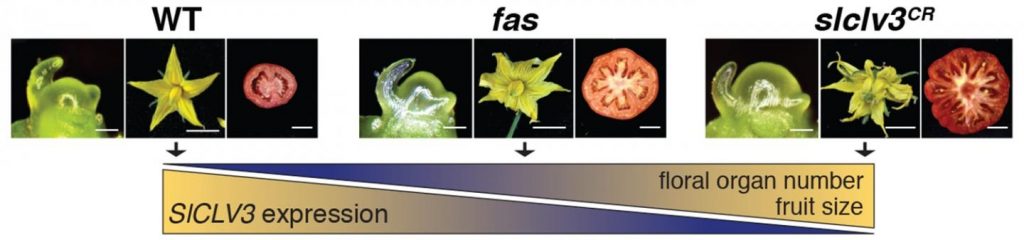 Using CRISPR to create different sets of mutations in a gene promoter called SICLV3 enabled researchers to vary the number of floral organs and locules in tomato
Using CRISPR to create different sets of mutations in a gene promoter called SICLV3 enabled researchers to vary the number of floral organs and locules in tomato