
Features
Agronomy
Genetics/Traits
New sources of clubroot resistance
Clubroot-resistant varieties are the key weapon for fighting this devastating disease in Prairie canola crops. However, experience in other countries shows the pathogen can evolve to overcome such resistance. The Canola Council of Canada’s announcement in April 2014 of the possible erosion of some forms of clubroot resistance in current canola varieties is a stark reminder that it can happen here, too.
Researchers in Saskatchewan are at work identifying additional resistance genes in canola’s cousins and transferring those genes into canola germplasm. Additional resistance genes can help in two ways: breeders can create cultivars with different resistance genes so growers can rotate between these different cultivars; and breeders can also pyramid the genes – put several different resistance genes together in the same cultivar. Both strategies make it harder for the pathogen to overcome the resistance. Also, the additional genes could protect against additional pathotypes, or strains, of the clubroot pathogen. The result will be more durable and broader-spectrum resistance to the pathogen.
Clubroot is caused by Plasmodiophora brassicae. This soil-borne pathogen causes irregular swellings (galls) on plant roots. The galls hinder the movement of water and nutrients into the top parts of the plant. Up to 100 per cent yield loss has occurred when susceptible canola cultivars were grown in clubroot-infested fields in Alberta.
In Prairie canola, clubroot was first found in a handful of fields in the Edmonton area in 2003. By 2013, surveys had confirmed a total of 1,482 infested fields in the province. The pathogen has also been found in Saskatchewan and Manitoba.
As the pathogen’s name suggests, Plasmodiophora brassicae affects canola and other species in the same family – Brassicaceae – including field crops like mustard, vegetables like cabbage, and weeds such as stinkweed.
One place to look for new sources of clubroot resistance is among these relatives of canola. That’s what Dr. Fengqun Yu, Dr. Gary Peng, Dr. Kevin Falk and several other researchers at Agriculture and Agri-Food Canada (AAFC) in Saskatoon have been working on, through a number of interrelated studies.
AAFC has provided the lion’s share of the funding for this work, but many other agencies have also given financial support. Some of these are the Saskatchewan Canola Development Commission (SaskCanola), Canola Council of Canada, Western Grains Research Foundation, Alberta Crop Industry Development Fund, Saskatchewan’s Agriculture Development Fund, Alberta Canola Producers Commission, and seed companies.
“Canola’s scientific name is Brassica napus. The sources of clubroot resistance in Brassica napus are very limited,” explains Yu. She says the resistance genes in the clubroot-resistant canola cultivars currently grown in Canada are from limited sources and many might have a connection with a single source: a European winter canola cultivar called Mendel.
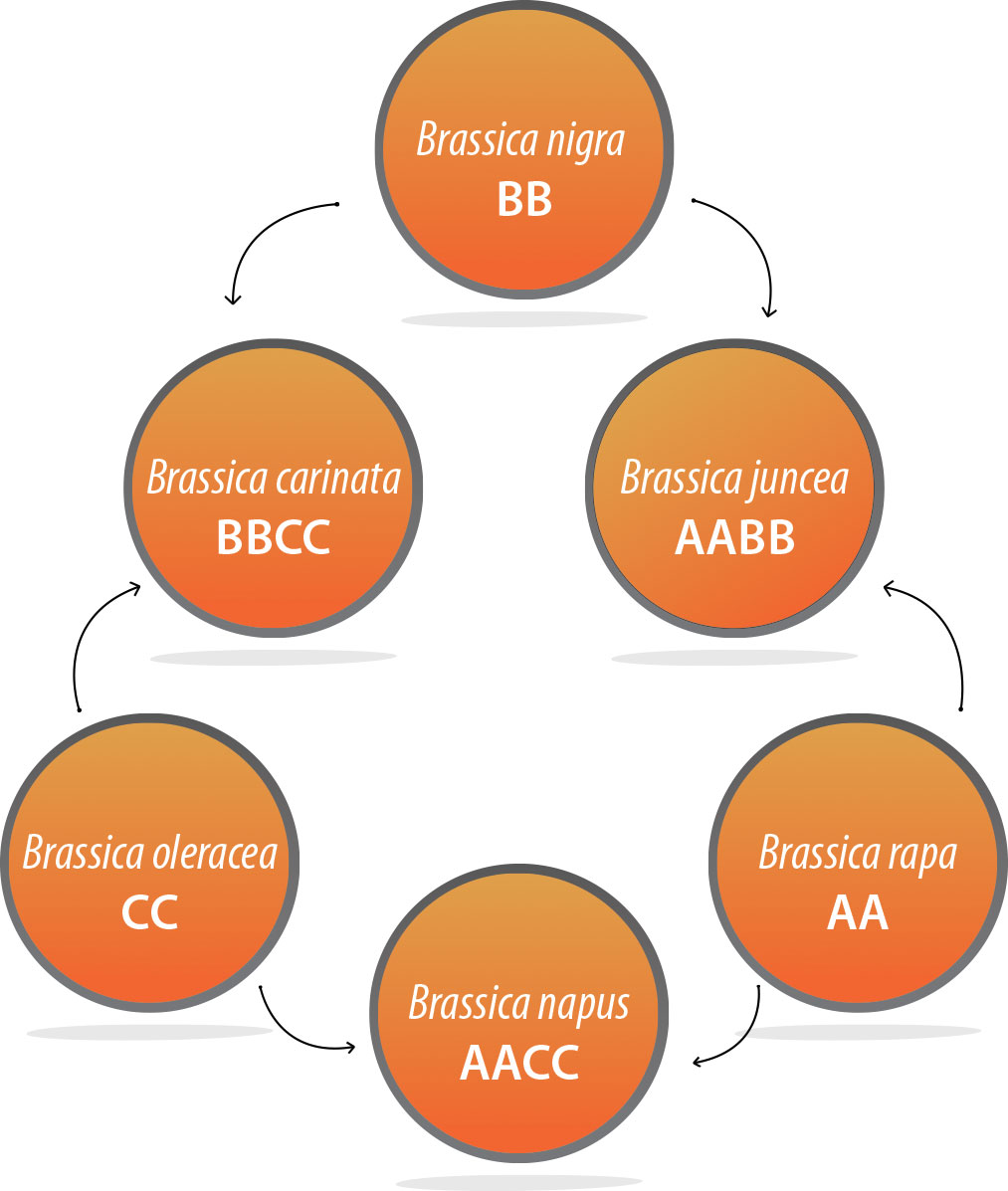
So the AAFC researchers are tapping into the genetic resources of six closely related Brassica species (see diagram below). Three of these are “diploids,” and three are “amphidiploids.” A diploid has two copies of each chromosome. An amphidiploid has two diploid chromosome sets, one from each parental species.
The genomes of the three diploids are referred to as AA, BB and CC. The AA genome is in Brassica rapa, which includes various subspecies such as Chinese cabbage and bok choy. Brassica nigra (black mustard) has the BB genome. And the CC genome is in Brassica oleracea, which has several subspecies such as cabbage, cauliflower and broccoli.
Brassica napus is an amphidiploid of Brassica rapa and Brassica oleracea, so it’s AACC. Brassica carinata (carinata or Ethiopian mustard) is BBCC, and Brassica juncea (brown mustard) is AABB.
Yu explains the reasons for seeking resistance genes in these Brassica species: “The first reason is that some of these diploid species, like Chinese cabbage, bok choy and cabbage, show very high levels of clubroot resistance against all pathotypes of the pathogen found in Canada.
“Secondly, transferring economically important traits from the diploid species B. rapa into canola is a well-established approach.” Yu herself has transferred three blackleg resistance genes from B. rapa into canola.
Peng and Falk led a study to screen close to 1,000 lines from the six species for clubroot resistance. They found 35 lines with resistance to pathotype 3 of the pathogen. Clubroot pathotypes are currently identified based on their ability to cause disease in a standardized set of hosts. In Alberta, the pathotypes include 2, 3, 5, 6 and 8. Pathotype 3, a very virulent type, is the most common pathotype in canola crops in the province.
Out of the 35 lines, the researchers decided to begin by working with eight very promising lines, most of which are B. rapa or B. oleracea. All eight have resistance against all five pathotypes.
Yu and several post-doctoral fellows are identifying resistance genes in these eight lines. In seven lines, they have found a single resistance gene, and in one line they have found two.
In all eight lines, they determined that clubroot resistance was a dominant trait. Yu explains, “Dominant resistance is very important for hybrid production. With dominant resistance, the hybrids just need resistance from one parent line. With recessive resistance, both parent lines would have to have the resistance gene.”
The researchers are also mapping the location of these genes within the chromosomes. To do this, they are using two methods that involve molecular markers. So far, they have mapped three of the genes: Rpb1 and Rpb2, which are both on the third chromosome in B. rapa; and Rpb3, on the eighth chromosome in B. rapa.
“For two of these genes, the markers are available already for breeders to use in marker-assisted selection. And for a third gene, the marker could be available within one year,” notes Yu.
The markers are being used in another component of this research: “introgressing” the resistance genes into canola germplasm. For example, to introgress resistance genes from B. rapa into B. napus, the researchers would cross a resistant B. rapa line with a B. napus breeding line and then identify the resistant offspring using the markers. Then they would backcross the resistant offspring with the B. napus line, identify the resistant offspring, and so on, to eventually develop germplasm very similar to the B. napus parent but with the resistance gene.
Rpb1, Rpb2 and Rpb3 are being introgressed into canola. A fourth clubroot resistance gene, Rpb4, identified in B. nigra, is being introgressed into B. carinata and B. juncea.
Yet another aspect of this research involves “resynthesizing” the amphidiploid species as a way to pyramid resistance genes. For instance, the researchers could cross a resistant B. rapa line and a resistant B. oleracea line to create a resynthesized B. napus line with resistance genes from each parent. They are resynthesizing B. napus, B. carinata and B. juncea and making good progress.
The results of all this work could soon contribute to new clubroot-resistant canola varieties.
“We are not in variety development; instead we develop clubroot-resistant canola germplasm that can be transferred to the seed companies,” explains Peng. “So far, we have transferred two sources of resistance to [a private company’s germplasm, under a collaborative agreement with that company], and we have transferred two resistance genes into our own breeding lines.” AAFC’s clubroot-resistant germplasm will be available to seed companies on a call-for-proposal basis.
In the meantime, Peng advises growers to do as much as they can to protect the resistance in the current clubroot-resistant canola varieties.
“Resistance is the cornerstone of clubroot management in Western Canada. Whenever possible, crop rotation and potentially cultivar rotation need to be kept in mind. Otherwise, eventually, the resistance will be eroded with the change in the pathogen population.”
 |
|
Relationships among Brassica species with the AA, BB and CC genomes, including canola (Brassica napus) |
November 6, 2014 By Carolyn King
 In these first-generation crosses between clubroot-resistant Brassica vegetables and a canola line Clubroot-resistant varieties are the key weapon for fighting this devastating disease in Prairie canola crops.
In these first-generation crosses between clubroot-resistant Brassica vegetables and a canola line Clubroot-resistant varieties are the key weapon for fighting this devastating disease in Prairie canola crops.