
Features
Agronomy
Plant Breeding
Mapping Phytophthora genome key to unravelling late blight
Potato breeders have a new tool to help develop potato resistance to the late blight pathogen Phytophthora infestans.
April 30, 2010 By Heather Hager
Potato breeders have a new tool to help develop potato resistance to the late blight pathogen Phytophthora infestans. Resistance has not yet proven durable because the pathogen evolves quickly, over-coming advances in potato genetics. However, scientists have now deciphered much of the P. infestans gene code, opening new avenues of research for resistance breeding.
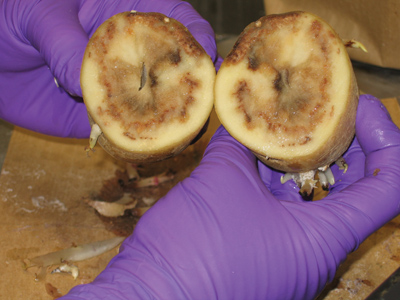 |
| Late blight was estimated in 2001 to cost the US potato industry $287.8 million in control costs and lost revenue, or $507 per hectare. Photo courtesy of Agnes Murphy, Potato Research Centre, Agriculture and Agri-Food Canada. |
Building on years of previous work, it took almost 100 scientists plus additional personnel from the US, Sweden, The Netherlands, the UK, Colombia, Israel and Germany several years to sequence and compile the P. infestans genome. The process involved sequencing many short, manageable lengths of the DNA and then assembling them in the correct order, like putting together a puzzle. Once the sequence was determined, computer programs compared it with known genes from similar organisms to give an idea of many of the genes’ functions.
Dr. Manuel Ospina-Giraldo, a molecular phytopathologist, and some of his undergraduate students at Lafayette College in Easton, Pennsylvania, were involved in sequencing the P. infestans genome. Ospina-Giraldo says that a couple of factors presented challenges to the sequencing process. For one, the P. infestans genome is very large compared to previously sequenced related species. At almost 250 million base pairs (bp), it is 2.6 times the size of the 95 million bp P. sojae, which causes soybean rot, and 3.8 times the size of the 65 million bp P. ramorum, which causes sudden oak death. In addition, the P. infestans genome has a large area that is composed of highly repetitive sequences. It is this latter area that is giving insight into the pathogen’s ability to overcome resistance.
It turns out that this highly repetitive region contains an unusually large number of genes that make effector proteins. “Effectors are something that the pathogen secretes, and they go into the plant cell and have an effect on the plant,” explains Dr. Richard Jones, a plant pathologist at the US Department of Agriculture Agricultural Research Service in Beltsville, Maryland. For example, some effectors from P. infestans might inhibit the plant’s ability to initiate a defence response against the pathogen. “P. infestans has a couple hundred of these, so it’s not clear what the function of each is,” says Jones, who also was involved in the sequencing project. Many of these effectors are under ongoing study.
Regions of the genome containing the effectors evolve rapidly, much more so than other regions containing genes that are required for everyday life functions. The pathogen’s general functioning is not greatly affected by mutations in the regions containing the effectors, so this genetic material can change rapidly without causing the organism’s death. Thus, these regions could be a key to understanding how the pathogen is able to evolve rapidly to keep up with advances in resistance.
In addition to effectors, some of P. infestans everyday genes might also contribute to its ability to infect potato plants. Because the pathogen has cell walls composed primarily of cellulose, similar to plants, Jones and Ospina-Giraldo think that the genes that produce enzymes involved in building and destroying its own cell walls may also play a role in destroying plant cell walls. “Penetration of the plant cell wall is the most critical aspect of the infection process,” says Ospina-Giraldo. “So penetration via the activity of these enzymes may aid infection.”
Close to 450 genes involved in cell wall synthesis and degradation have been identified in P. infestans.
Breeding a resistant potato
“Breeding for resistance against P. infestans is really high in priority with regard to our objectives because it’s a big issue,” says Dr. Benoit Bizimungu, potato breeder at Agriculture and Agri-Food Canada’s Potato Research Centre in Fredericton, New Brunswick. Chemical control measures are costly, as can be potato losses due to the pathogen. Then there are potential environmental and health effects of frequent chemical applications, as well as the development of more aggressive pathogen populations and increasing pathogen resistance to the chemicals. And there are negative effects on society through potential potato losses leading to decreased food availability.
 |
| Potato plants are screened for resistance to the leaf blight pathogen under artificial inoculation in the laboratory. Photo courtesy of Agnes Murphy, Potato Research Centre, Agriculture and Agri-Food Canada.
|
Breeding for resistance has been difficult because the pathogen mutates and adapts so quickly. Now, knowing the pathogen’s genome will allow a better understanding of how it infects the plant and how it can overcome resistance genes, which should help breeders to develop more durable resistance, says Bizimungu. He notes that the potato genome is also being sequenced. Combining the information from both genomes should speed up the development of new varieties, he says. “In any disease, you need to know the pathogen first in order to develop ways to attack or control the pathogen,” says Ospina-Giraldo. By knowing how the pathogen manipulates the plant to promote infection, researchers might be able to develop plants that block those processes. This could be done through traditional plant breeding, perhaps using molecular markers to pinpoint specific genes. Another option is through genetic engineering to develop plants that produce specific inhibitors against infection processes.
For example, assuming that certain cell-wall-degrading enzymes are important in infection, Jones is looking at genes and processes in the potato plant that inhibit those enzymes. Those could then be enhanced in the plant. “We work on both the P. infestans side and the potato side,” he says. “With both genomes sequenced, it will help to make matches between genes in the pathogen and potential counter-genes in the potato.”
The question is: how long will it take to make a better potato? Through its regular breeding program, AAFC already has some new, improved potato selections that are under evaluation, says Bizimungu. Any advances in durable resistance breeding that are gained using the new genome information “will not come out tomorrow,” he says, but the improvement in the knowledge of the pathogen, and of the potato plant, should speed up the development of new varieties with better resistance to the late blight pathogen.
| Phytophthora is not a fungus Although commonly mistaken as one, Phytophthora is not a fungus. In fact, whereas fungi are more closely related to animals, Phytophthora is more closely related to plants, in particular, diatoms (single-celled algae) and brown algae (seaweeds), explains Dr. Manuel Ospina-Giraldo, a molecular phytopathologist at Lafayette College in Easton, Pennsylvania. Additionally, the cell walls of fungi are made primarily of chitin, whereas those of Phytophthora are made of cellulose, similar to the cell walls of plants. |